Example: Introduction to topsbm¶
Topic modelling with hierarchical stochastic block models
[1]:
from sklearn.feature_extraction.text import CountVectorizer
import pandas as pd
from topsbm import TopSBM
Setup: Load a corpus¶
- We have a list of documents, each document contains a list of words.
- We have a list of document titles (optional)
The example corpus consists of 63 articles from Wikipedia taken from 3 different categories (Experimental Physics, Chemical Physics, and Computational Biology).
We use scikit-learn’s CountVectorizer to turn this text into a feature matrix.
[2]:
# Load texts and vectorize
with open('corpus.txt', 'r') as f:
docs = f.readlines()
vec = CountVectorizer(token_pattern=r'\S+')
X = vec.fit_transform(docs)
# X is now a sparse matrix of (docs, words)
# titles corresponding to docs
with open('titles.txt', 'r') as f:
x = f.readlines()
titles = [h.split()[0] for h in x]
[3]:
# view the data for document 0
print(titles[0])
print(docs[0][:100])
Nuclear_Overhauser_effect
the nuclear overhauser effect noe is the transfer of nuclear spin polarization from one nuclear spi
Fit the model¶
Calling TopSBM.fit_transform will: * construct the bipartite graph between documents and words (samples and features) * perform Hierarchical Stochastic Block Model inference over the graph * return an embedding of the samples in the block level with finest granularity
[18]:
model = TopSBM(random_state=9)
Xt = model.fit_transform(X)
Plotting the graph and block structure¶
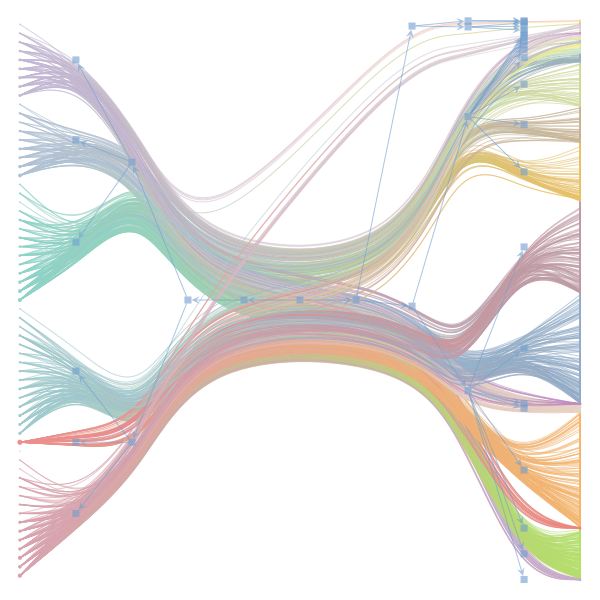
The following plot shows the (hierarchical) community structure in the word-document network as inferred by the stochastic block model:
- document-nodes are on the left
- word-nodes are on the right
- different colors correspond to the different groups
The result is a grouping of nodes into groups on multiple levels in the hierarchy:
- on the uppermost level, each node belongs to the same group (square in the middle)
- on the next-lower level, we split the network into two groups: the word-nodes and the document-nodes (blue sqaures to the left and right, respectively). This is a trivial structure due to the bipartite character of the network.
- only next lower levels constitute a non-trivial structure: We now further divide nodes into smaller groups (document-nodes into document-groups on the left and word-nodes into word-groups on the right)
In the code, the lowest level is known as level 0, with coarser levels 1, 2, …
[19]:
model.plot_graph(n_edges=1000)

Topics¶
For each word-group on a given level in the hierarchy, we retrieve the 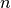 most common words in each group – these are the topics!
most common words in each group – these are the topics!
[20]:
topics = pd.DataFrame(model.groups_[1]['p_w_tw'],
index=vec.get_feature_names())
[21]:
for topic in topics.columns:
print(topics[topic].nlargest(10))
print()
the 0.006768
of 0.006661
a 0.006554
in 0.006446
is 0.006446
to 0.006339
and 0.006124
for 0.005372
an 0.005264
as 0.005264
Name: 0, dtype: float64
when 0.008921
where 0.008058
first 0.006619
given 0.006331
if 0.006331
field 0.006043
applied 0.005755
because 0.005755
e 0.005468
energy 0.005468
Name: 1, dtype: float64
computational 0.060606
development 0.055556
proteins 0.045455
open 0.040404
protein 0.040404
software 0.040404
community 0.035354
researchers 0.035354
core 0.025253
identify 0.025253
Name: 2, dtype: float64
point 0.217391
formula 0.188406
must 0.144928
wave 0.115942
spectrum 0.086957
air 0.072464
plane 0.057971
flow 0.043478
q 0.043478
mode 0.028986
Name: 3, dtype: float64
Topic-distribution in each document¶
Which level-1 topics contribute to each document?
[22]:
pd.DataFrame(model.groups_[1]['p_tw_d'],
columns=titles)
[22]:
| Nuclear_Overhauser_effect | Quantum_solvent | Rovibrational_coupling | Effective_field_theory | Chemical_physics | Rotational_transition | Dynamic_nuclear_polarisation | Knight_shift | Polarizability | Anisotropic_liquid | ... | Louis_and_Beatrice_Laufer_Center_for_Physical_and_Quantitative_Biology | Law_of_Maximum | Enzyme_Function_Initiative | SnoRNA_prediction_software | Sepp_Hochreiter | Aureus_Sciences | IEEE/ACM_Transactions_on_Computational_Biology_and_Bioinformatics | Knotted_protein | BioUML | De_novo_transcriptome_assembly | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.608392 | 0.856 | 0.523529 | 0.804651 | 0.82 | 0.648649 | 0.584192 | 0.582418 | 0.493274 | 0.645161 | ... | 0.907692 | 0.851351 | 0.857143 | 0.857143 | 0.846690 | 0.822222 | 0.84375 | 0.773585 | 0.870647 | 0.868932 |
| 1 | 0.391608 | 0.144 | 0.458824 | 0.190698 | 0.16 | 0.337838 | 0.412371 | 0.406593 | 0.500000 | 0.354839 | ... | 0.092308 | 0.121622 | 0.095238 | 0.142857 | 0.139373 | 0.133333 | 0.09375 | 0.169811 | 0.084577 | 0.092233 |
| 2 | 0.000000 | 0.000 | 0.000000 | 0.000000 | 0.00 | 0.000000 | 0.000000 | 0.010989 | 0.002242 | 0.000000 | ... | 0.000000 | 0.027027 | 0.044218 | 0.000000 | 0.013937 | 0.044444 | 0.06250 | 0.047170 | 0.044776 | 0.038835 |
| 3 | 0.000000 | 0.000 | 0.017647 | 0.004651 | 0.02 | 0.013514 | 0.003436 | 0.000000 | 0.004484 | 0.000000 | ... | 0.000000 | 0.000000 | 0.003401 | 0.000000 | 0.000000 | 0.000000 | 0.00000 | 0.009434 | 0.000000 | 0.000000 |
4 rows × 63 columns
Extra: Clustering of documents - for free.¶
The stochastic block models clusters the documents into groups. We do not need to run an additional clustering to obtain this grouping.
For a query article, we can return all articles from the same group
[23]:
cluster_labels = pd.DataFrame(model.groups_[1]['p_td_d'],
columns=titles).idxmax(axis=0)
cluster_idx = cluster_labels['Rovibrational_coupling']
cluster_labels[cluster_labels == cluster_idx]
[23]:
Nuclear_Overhauser_effect 0
Rovibrational_coupling 0
Rotational_transition 0
Dynamic_nuclear_polarisation 0
Knight_shift 0
Polarizability 0
Anisotropic_liquid 0
Rotating_wave_approximation 0
Molecular_vibration 0
Fuel_mass_fraction 0
Electrostatic_deflection_(structural_element) 0
Magic_angle_(EELS) 0
Reactive_empirical_bond_order 0
Photofragment-ion_imaging 0
Molecular_beam 0
McConnell_equation 0
Ziff-Gulari-Barshad_model 0
Empirical_formula 0
Newton's_laws_of_motion 0
Ripple_tank 0
Particle-induced_X-ray_emission 0
Elevator_paradox_(physics) 0
Wave_tank 0
X-ray_crystal_truncation_rod 0
Faraday_cup_electrometer 0
Line_source 0
X-ray_standing_waves 0
Point_source 0
Fragment_separator 0
Dynamic_mode_decomposition 0
Euler's_laws_of_motion 0
Quantum_oscillations_(experimental_technique) 0
dtype: int64